Human Imprintome Genomic Map
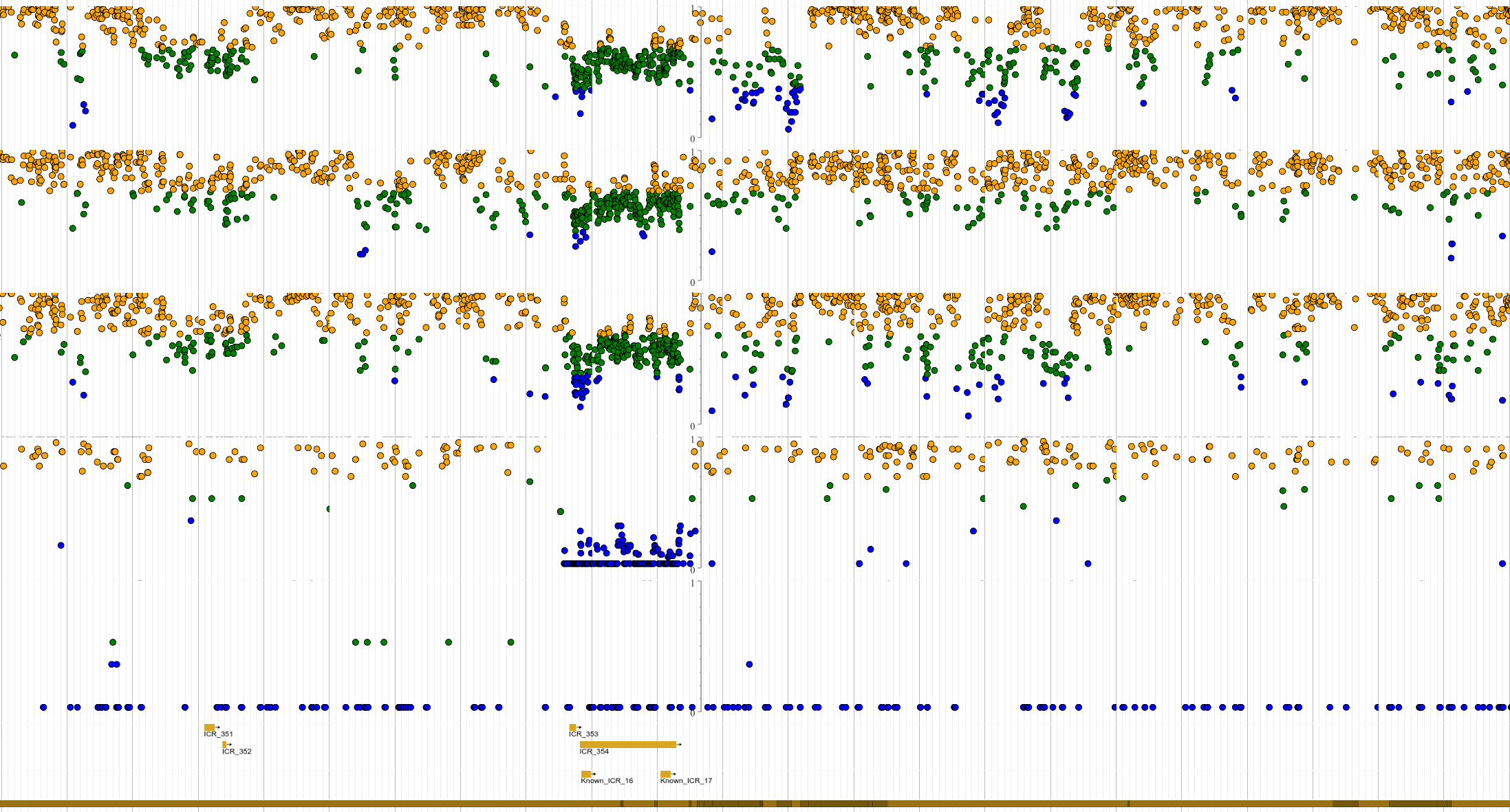
A new paper from CHHE members and students Dereje D. Jima, David A. Skaar, Antonio Planchart, Alison Motsinger-Reif, Sebnem E. Cevik, Sarah S. Park, Michael Cowley, Fred Wright, John House, Andy Liu, Randy L. Jirtle, and Cathrine Hoyo published in the journal Epigenetics highlights human imprint genes and provides an opportunity to evaluate environmental epigenetic impacts.
Genomic imprinting is a unique epigenetic form of gene regulation that evolved in
marsupials and placental mammals about 150 million years ago. It results in only one
copy of a gene being expressed in a parent-of-origin dependent manner. Thus,
imprinted genes are disease susceptibility loci since a single genetic or epigenetic event
can alter their function.
Imprinted genes are heavily involved in metabolism and growth regulation. As a
consequence, an epigenetic tug-of-war between the mother’s and father’s imprinted
genes during development may explain variations in the fetal origins of metabolic
disorders such as obesity and type 2 diabetes, as well as cancer, and a spectrum of
mental disorders from autism to schizophrenia.
Using whole genome bisulfite sequencing, our investigation identified 1,488
hemimethylated DNA candidate imprint control regions (ICRs) in the human genome
that potentially regulate parental-specific expression of imprinted genes – the human
imprintome (https://doi.org/10.1080/15592294.2022.2091815). As a consequence, it is
now possible to investigate the role of environmentally-induced epigenetic changes in
ICRs during early development in the pathogenesis of human diseases and behavioral
disorders. Furthermore, since the ICRs are established before tissue differentiation,
these studies can be performed with cell samples readily available to epidemiologists
(e.g., white blood cells and buccal epithelial cells), greatly facilitating environmental
epigenomics studies.
Imprintome Website: https://humanicr.org/
- Categories: